Evolution Weasel Program
HAMLET: Do you see yonder cloud that’s almost in shape of a camel?
POLONIUS: By th' mass, and ’tis like a camel indeed.
HAMLET: Methinks it is like a weasel.
POLONIUS: It is backed like a weasel.
HAMLET: Or like a whale.
POLONIUS: Very like a whale.

The weasel program, Dawkins' weasel, or the Dawkins weasel is a thought experiment and a variety of computer simulations illustrating it. Their aim is to demonstrate that the process that drives evolutionary systems—random variation combined with non-random cumulative selection—is different from pure chance.
Weasel program - Wikipedia
The thought experiment was formulated by Richard Dawkins, and the first simulation written by him; various other implementations of the program have been written by others.
The Weasel algorithm implemented here runs as follows:
- Start - A random state (a string of 28 characters).
- Produce offspring - Make N copies of the string.
- Mutate - For each character in each of the copies, with a probability of P, replace the character with a new random character.
- Compare each new string with the target string
METHINKS IT IS LIKE A WEASEL, and give each a score.- The number of offspring N and the probability P are configurable (default values: N=100 and P=5%)
- The algorithm used for scoring is Hamming distance
- In real life there is no final pre-established target!
- Survival - Take the highest scoring string, and go to step 2.
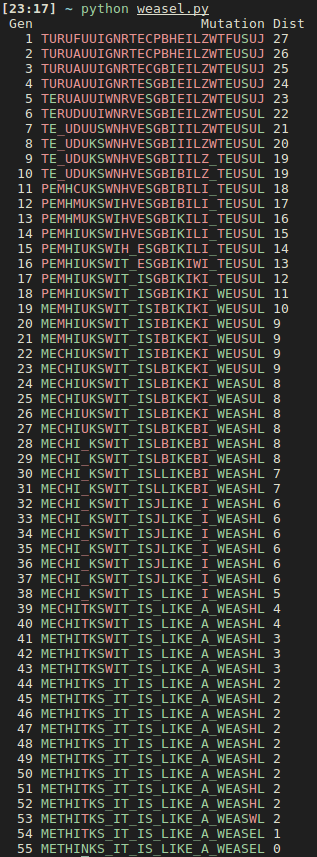
Show Me The Code
import string
import random
import itertools
# Possible genes (whole character set)
charset = string.ascii_uppercase + "_"
# Number of genes (characters in the string)
numGenes = 28
# Create a next generation of mutated offspring
def mutate(state, numOffspring, mutationProb):
mutations = []
for i in range(numOffspring):
mutation = ""
for j in range(numGenes):
# Will character j be mutated?
p = random.randint(1, 100)
if p<=mutationProb:
newGene = charset[random.randint(0, len(charset)-1)]
mutation += newGene
else:
mutation += state[j]
mutations.append(mutation)
return mutations
# Compute Hamming distance between two strings
def hamming(str1, str2):
return sum(itertools.imap(str.__ne__, str1, str2))
# Select the fittest offspring from the pool
# (Live free or die - UNIX)
def fittest(mutations, target):
min = numGenes+1
fittest = None
for m in mutations:
d = hamming(m, target)
if d<min:
min = d
fittest = m
return fittest
# Colourise mutation based on distance from target
# (We don't care about performance so light the Christmas tree)
def colorise(mutation, target):
W = '\033[0m' # white (normal)
R = '\033[31m' # red
G = '\033[32m' # green
s = ""
for i in range(len(mutation)):
if mutation[i] == target[i]:
s += G + mutation[i]
else:
s += R + mutation[i]
s += W
return s
# While target not reached, evolve!
def evolve(numOffspring, mutationProb):
cur = "".join(random.choice(charset) for _ in range(numGenes))
fin = "METHINKS_IT_IS_LIKE_A_WEASEL"
gen = 0
print "%4s %28s %4s" % ("Gen", "Mutation", "Dist")
while cur != fin:
offspring = mutate(cur, numOffspring, mutationProb)
cur = fittest(offspring, fin)
gen += 1
print "%4d" % gen, colorise(cur, fin), hamming(cur, fin)
if __name__ == "__main__":
# Number of offspring per generation
numOffspring = 100
# Probability that a gene (character) will mutate (percent)
mutationProb = 5
evolve(numOffspring, mutationProb)
Conclusions
- It’s interesting to vary the number of offsping per generation and the mutation probability and see how generations evolve:
- For a higher number of offspring N the generations evolve much more quickly towards the target.
- For a higher mutation probability P evolution becomes random!